Influenza Genome Sequencing Project
This article needs additional citations for verification. (November 2007) |
| Influenza (flu) |
|---|
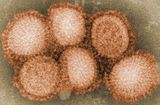 |
The Influenza Genome Sequencing Project (IGSP), initiated in early 2004, seeks to investigate influenza evolution by providing a public data set of complete influenza genome sequences from collections of isolates representing diverse species distributions.
The project is funded by the National Institute of Allergy and Infectious Diseases (NIAID), a division of the National Institutes of Health (NIH), and has been operating out of the NIAID Microbial Sequencing Center at The Institute for Genomic Research (TIGR, which in 2006 became The Venter Institute). Sequence information generated by the project has been continually placed into the public domain through GenBank.
Origins[]
In late 2003, David Lipman, , Steven Salzberg, and a consortium of other scientists wrote a proposal to begin sequencing large numbers of influenza viruses at The Institute for Genomic Research (TIGR). Prior to this project, only a handful of flu genomes were publicly available.[citation needed] Their proposal was approved by the National Institutes of Health (NIH), and would later become the IGSP. New technology development led by Elodie Ghedin began at TIGR later that year, and the first publication describing > 100 influenza genomes appeared in 2005 in the journal Nature [1]
Research goals[]
The project makes all sequence data publicly available through GenBank, an international, NIH-funded, searchable online database. This research helps to provide international researchers with the information needed to develop new vaccines, therapies and diagnostics, as well as improve understanding of the overall molecular evolution of Influenza and other genetic factors that determine their virulence.[citation needed] Such knowledge could not only help mitigate the impact of annual influenza epidemics, but could also improve scientific knowledge of the emergence of pandemic influenza viruses.
Results[]
The project completed its first genomes in March 2005 and has rapidly accelerated since. By mid-2008, over 3000 isolates had been completely sequenced from influenza viruses that are endemic in human ("human flu") avian ("bird flu") and swine ("swine flu") populations, including many strains of H3N2 (human), H1N1 (human), and H5N1 (avian).[1]
Affiliations[]
The project is funded by the National Institute of Allergy and Infectious Diseases (NIAID) which is a component of the NIH, which is an agency of the United States Department of Health and Human Services.
The IGSP has expanded to include a growing list of collaborators, who have contributed both expertise and valuable collections of influenza isolates. Key early contributors included Peter Palese of the Mount Sinai School of Medicine in New York, Jill Taylor of the Wadsworth Center at the New York State Department of Health, Lance Jennings of (New Zealand), Jeff Taubenberger of the Armed Forces Institute of Pathology (who later moved to NIH), Richard Slemons of Ohio State University and Rob Webster of St. Jude's Children's Hospital in Memphis, Tennessee.
In 2006 the project was joined by Ilaria Capua of the (in Italy), who contributed a valuable collection of avian flu isolates (including multiple H5N1 strains). Some of these avian isolates were described in a publication in Emerging Infectious Diseases in 2007.[2] Nancy Cox from the Centers for Disease Control and Prevention (CDC) and Robert Couch from Baylor College of Medicine also joined the project in 2006, contributing over 150 influenza B isolates.
The project began prospective studies of the 2007 influenza season with collaborators Florence Bourgeois and Kenneth Mandl of Children's Hospital Boston and the Harvard School of Public Health and Laurel Edelman of Surveillance Data Inc.[citation needed]
References[]
- ^ Jump up to: a b Ghedin E, Sengamalay NA, Shumway M, Zaborsky J, Feldblyum T, Subbu V, Spiro DJ, Sitz J, Koo H, Bolotov P, Dernovoy D, Tatusova T, Bao Y, St George K, Taylor J, Lipman DJ, Fraser CM, Taubenberger JK, Salzberg SL (October 2005). "Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution". Nature. 437 (7062): 1162–6. Bibcode:2005Natur.437.1162G. doi:10.1038/nature04239. PMID 16208317. [More Flu Viruses Around than Expected, Mutate in Unexpected Ways Lay summary] Check
|lay-url=value (help). (as PDF - ^ Salzberg SL, Kingsford C, Cattoli G, et al. (May 2007). "Genome analysis linking recent European and African influenza (H5N1) viruses". Emerging Infect. Dis. 13 (5): 713–8. doi:10.3201/eid1305.070013. PMC 2432181. PMID 17553249.
External links[]
- Influenza Sequencing Project home page at TIGR
- Influenza Genome Sequencing Project home page at NIAID
- Influenza virus resource at NCBI (NIH)
- Fauci AS (January 2006). "Pandemic influenza threat and preparedness". Emerging Infect. Dis. 12 (1): 73–7. doi:10.3201/eid1201.050983. PMC 3291399. PMID 16494721.
- Influenza Research Database Database of influenza sequences and related information.
- National Institutes of Health
- Genome projects
- Influenza
